 
The algorithm is based on sequencing reads alignment with reference V gene and J gene segments for the identification of TCR sequences, as well as applying a 96% cutoff value on J gene segment diversity. More recently, the iSSAKE-like strategy was developed by Warren et al. Because of the sequencing cost, short paired-end reads generated by next-generation sequencing technology are and will remain the main source of data, and we believe that the TCRklass is a useful and reliable toolkit for TCR repertoire analysis. 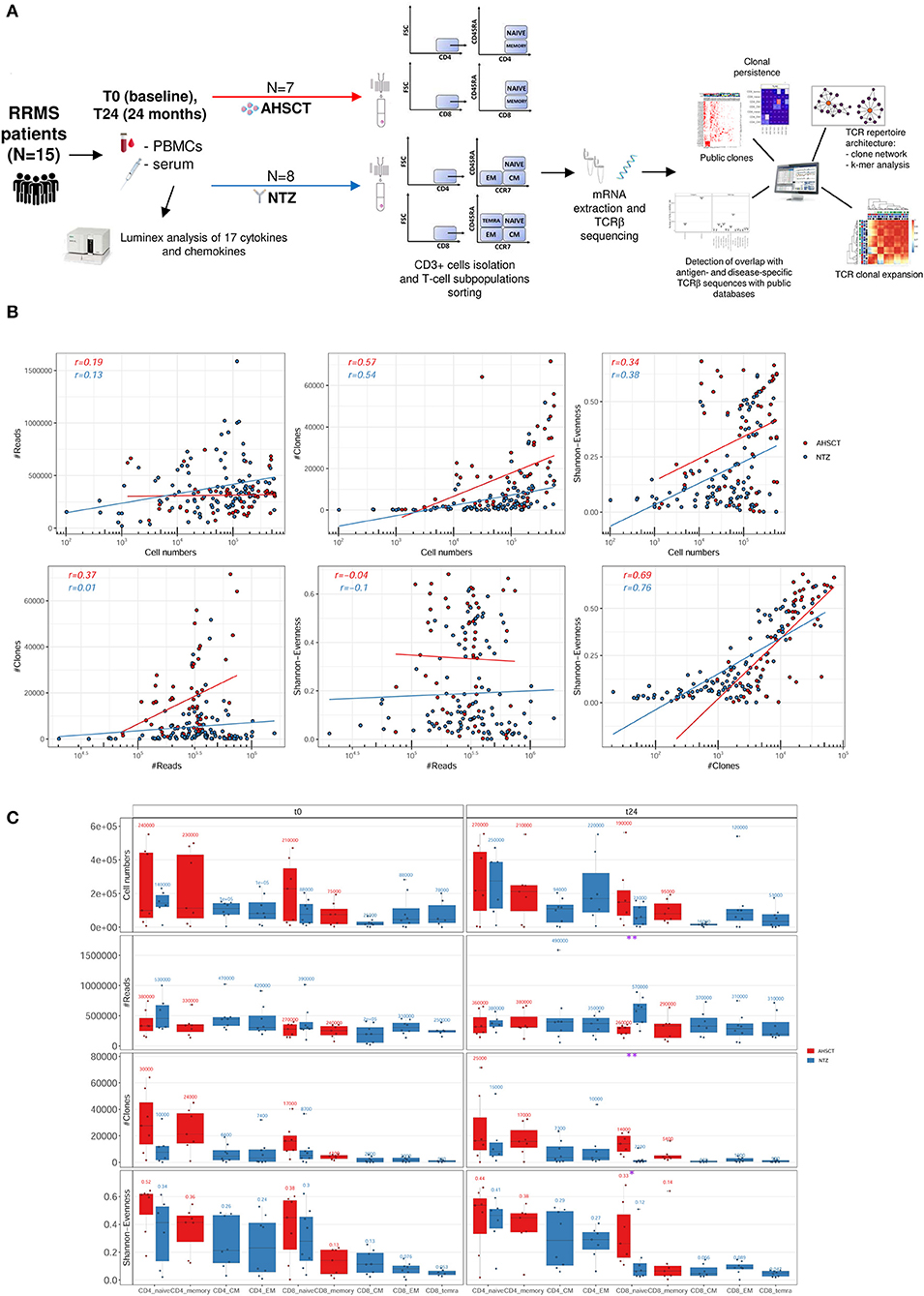
We applied TCRklass on large datasets of two human and three mouse TCR repertoires it demonstrated higher reliability on CDR3 identification and much less biased V/J profiling, which are the two components contributing the diversity of the repertoire. We tested TCRklass using manually curated short read datasets in comparison with in silico datasets it showed higher precision and recall rates on CDR3 identification. To decipher the complexity of TCR repertoire, we developed an integrated pipeline, TCRklass, using K-string–based algorithm that has significantly improved the accuracy and performance over existing tools. Genome Res 2011 21:790–797.The next-generation sequencing technology has promoted the study on human TCR repertoire, which is essential for the adaptive immunity. Exhaustive T-cell repertoire sequencing of human peripheral blood samples reveals signatures of antigen selection and a directly measured repertoire size of at least 1 million clonotypes. Next-generation-sequencing-spectratyping reveals public T-cell receptor repertoires in pediatric very severe aplastic anemia and identifies a β chain CDR3 sequence associated with hepatitis-induced pathogenesis. Overlap and effective size of the human CD8+ T cell receptor repertoire. Robins HS, Srivastava SK, Campregher PV, et al. Profiling the T-cell receptor beta-chain repertoire by massively parallel sequencing. Unraveling V(D)J recombination: insights into gene regulation. Moreover, our data show that terminal deoxynucleotidyl transferase activity was biased toward the insertion of G (31.92%) and C (27.14%) over A (21.82%) and T (19.12%) nucleotides.Some conserved features could be observed in the composition of CDR3, which may inform future studies of human TCR gene recombination. Notably, the usage frequency of individual nucleotides and amino acids within complementarity-determining region (CDR3) intervals was remarkably consistent between individuals. The usage frequencies of the TCR beta variable, beta joining, and beta diversity gene segments were similar among T cells from different individuals. We then combined multiplex-PCR, Illumina sequencing, and IMGT/High V-QUEST to analyze the characteristics and polymorphisms of the TCR.Most of the individual T cell clones were present at very low frequencies, suggesting that they had not undergone clonal expansion. T cells were isolated with anti-human CD3 magnetic beads according to the manufacturer's protocol. Herein, we used this technology to study the repertoire features of TCR beta chain in the blood of healthy individuals.Peripheral blood samples were collected from 10 healthy donors. 
Next-generation sequencing has become a powerful tool for deep TCR profiling. The T-cell receptor (TCR) repertoire is a mirror of the human immune system that reflects processes caused by infections, cancer, autoimmunity, and aging. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
